not in multiples of 3) gaps may be inserted, since the alignment program often does not take theĬoding nature of the sequence into account. When coding nucleotide sequences are aligned directly,įrameshifting (i.e. Your alignment should exclude any non-coding region of the nucleotide sequence, such as introns or promoter regions,įor which existing models of codon substitution would not apply. due to misalignment, or a non-functional coding sequence) and the terminal stop codon. that it does not contain stop codons, including premature stop codons You should verify that the alignment is in frame, i.e. Then it will not be properly read by Datamonkey.
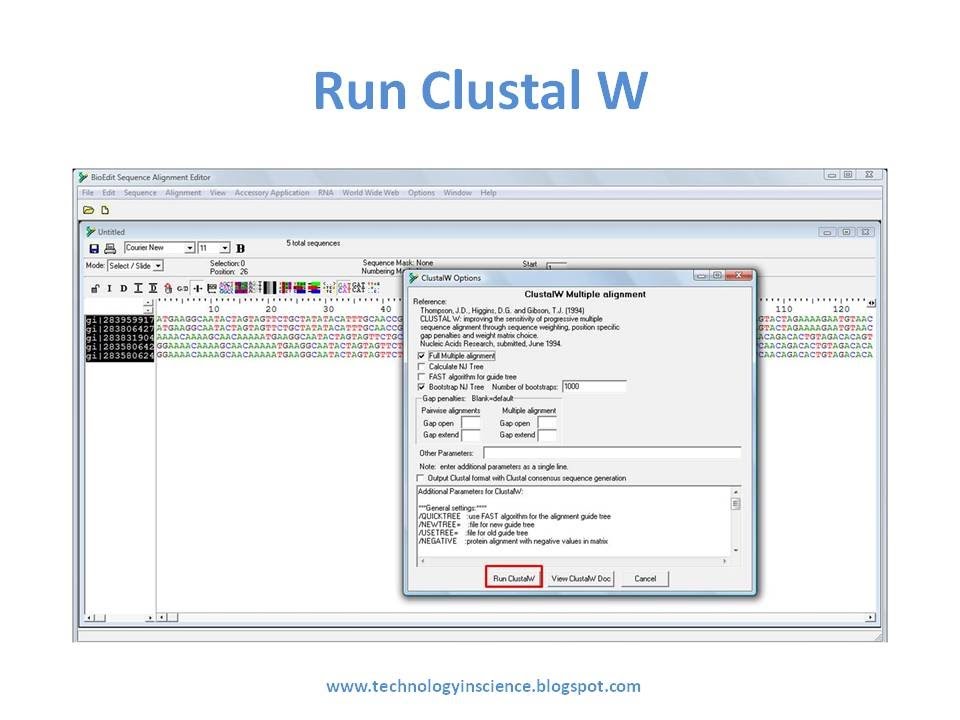
Align in bioedit software#
sequences that are out of frame)Īre easy to spot with software that provides a graphical visualization of the alignment, such as HyPhy, Se-Al, or BioEdit.ĭatamonkey uses the HyPhy package as its processing engine, and if an alignment does not open in HyPhy on your machine (using the File:Open:Open Data File command), Of course, one can never be sure that an alignment is objectively ÒcorrectÓ, but gross misalignments (e.g. It is a good practice to visually inspect your data to make sure that the sequences are alignment correctly. Selection using HyPhy), and we refer an interested reader to A number of publications have dealt with this issue extensively (e.g. Selective sweep are included in the sample. (rapid replacement of one allele with a more fit one, resulting in a homogeneous population), unless sequences sampled prior to and following the For example, they should not be applied to the detection of selective sweeps The median number of sequences in an alignment submitted to Datamonkey is 19.Ĭomparative methods are ill suited to study certain kinds of selection. While as few as 4 may be sufficient for alignment-wide inference (PARRIS/GA-Branch).

The number of sequences in the alignment is important: too few sequences will contain too little information for meaningful inference, while too many may take too long to run.Īt the time of this writing, Datamonkey permits up to 150 sequences for SLAC analyses, 100 for FEL/IFEL analyses, 40 for REL and PARRIS and 25 for GA-Branch.Īs a rule of thumb, at least 10 sequences are needed to detect selection at a single site (SLAC/FEL/IFEL/REL) with any degree of reliability, our inability to reliably infer branch lengths and substitution parameters. However, sequences that are too divergent could lead to saturation, i.e. Phylogenetic tree should be at least one expected substitution per codon site, but it is impossible to give a generally valid range for desirable sequence divergence. Yang and colleagues have suggested that the total length of the Sequence along a star phylogeny) sample of the Human T-lymphotropic virus (HTLV), they found that the method performed poorly. For example when, Suzuki and Nei applied a REL-type method to a very low divergence (1 or 2 substitutions per Sequence diversity is needed for reliable inference.
Because comparative methods estimate relative rates of synonymous and non-synonymous substitution, substantial Influenza A viruses infecting different individuals). mammalian interferon genes), or a diverse population sample Ideally, the alignment should represent a single gene, or protein product, sampled over multiple taxa (e.g. To perform a selection analysis, needs a multiple alignment of at least three homologous coding nucleotide sequences.Ĭodon based methods for estimating dN and dS can be applied to any sequence alignment, but there are several considerations to keep in mind:
